Volume 6, Issue 2 (Journal of Clinical and Basic Research (JCBR) 2022)
jcbr 2022, 6(2): 1-11 |
Back to browse issues page
Download citation:
BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:



BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:
Rostami M, Mirarab A, Mohebbi A. Quaking Gene Expression as a Prognostic Marker in Neural Inflammation Disorders. jcbr 2022; 6 (2) :1-11
URL: http://jcbr.goums.ac.ir/article-1-354-en.html
URL: http://jcbr.goums.ac.ir/article-1-354-en.html
1- Department of Immunotherapy and Vaccine Research, Pasteur Institute of Iran, Tehran, Iran
2- Department of Microbiology, School of Medicine, Golestan University of Medical Sciences, Gorgan, Iran
3- Vista Aria Rena Gene Inc., Golestan Province, Gorgan, Iran ,Mohebbi-a@goums.ac.ir
2- Department of Microbiology, School of Medicine, Golestan University of Medical Sciences, Gorgan, Iran
3- Vista Aria Rena Gene Inc., Golestan Province, Gorgan, Iran ,
Full-Text [PDF 712 kb]
(1201 Downloads)
| Abstract (HTML) (2909 Views)
Full-Text: (912 Views)
INTRODUCTION
The extracellular matrix, mesenchymal stem cells, cancer-associated fibroblasts, endothelial cells, immune cells, and a complex network of cytokines and growth factors make up the tumor microenvironment (1). Tumor tissues are composed of a variety of cancer cells, including cancer stem cells (CSCs), which can differentiate into cancer cells (2,3). The stem cell niche is a milieu in which tissue-specific stem cells in normal organs maintain their stemness. The most common and deadly primary brain tumor is glioblastoma (GBM). Glioma stem cells are thought to have a role in tumors' resistance to standard therapy. These cells have distinct surface markers, control specific signaling pathways, and play an important role in the development of glioma vessels (4,5). Niches have been identified in multiple cancers, and they often assimilate the signals of niches for the tissues from which the CSCs appeared (6,8). However, it is not understood how these cells still manage to preserve their nature when they invade and migrate from their homes to other areas where ideal niches are less likely to be available (9).
Brain tumors and metastases represent a heterogeneous set of conditions (10). Primary brain tumors such as astrocytic tumors, oligodendrogliomas, ependymomas, and mixed gliomas are all referred to as gliomas (11). Adults' most usual lethal brain tumor is grade IV glioma also known as GBM, with over 10,000 cases diagnosed annually in the United States (12). Evidence shows that a population of GBM cells has a stable ability to self-renew and produce new tumors that keep the features of original tumors (13,14).
The ability of stem cells to invade and migrate raises the question of how these cells maintain their stemness when they
come into contact with varied ingredient compositions of a new microenvironment (15). Therefore, looking for genetic changes could help CSCs to keep stemness outside their niches (16). The Quaking gene (QKI), as a tumor suppressor gene, may poten¬tially affect CSC stemness. The gene encodes QKI, a STAR-family RNA-binding protein that is involved in RNA homeostasis (19,20). The RNA-binding protein QKI is highly expresses in brain cells (17), and play a regulatory role in brain development via modulating the stability of mRNAs that promote differentiation and inhibiting cell cycle progress (18). In the present study, the significance of QKI mRNA in human central nervous system (CNS) cancer was evaluated by using the Cancer Genome Atlas (TCGA) data portals.
MATERIALS AND METHODS
Analysis for QKI mutations
The Catalog of Somatic Mutations in Cancer (COSMIC) database was used to analyze QKI mutations (21). Pie charts were generated for a distribution survey and substitutions on the coding strand in CNS cancer.
CBioPortal analysis for alteration frequency of QKI
Alteration frequency of QKI mRNA was performed using cBioPortal for Cancer Genomics (22,23). All searches were performed according to the cBioPortal’s online instructions. The database query was based on mutation and altered expression of the QKI in six CNS/brain studies (Brain Low Grade Glioma (TCGA, Firehose Legacy), Brain Low Grade Glioma (TCGA PanCancer Atlas), GBM (TCGA, Cell 2013), GBM (TCGA, Nature 2008), Low-Grade Gliomas (UCSF, Science 2014), Merged Cohort of LCG and GBM (TCGA, Cell 2016), including 3010 samples of 2958 patients (summarized in Figure 1). The overlapped studies were discarded.

The extracellular matrix, mesenchymal stem cells, cancer-associated fibroblasts, endothelial cells, immune cells, and a complex network of cytokines and growth factors make up the tumor microenvironment (1). Tumor tissues are composed of a variety of cancer cells, including cancer stem cells (CSCs), which can differentiate into cancer cells (2,3). The stem cell niche is a milieu in which tissue-specific stem cells in normal organs maintain their stemness. The most common and deadly primary brain tumor is glioblastoma (GBM). Glioma stem cells are thought to have a role in tumors' resistance to standard therapy. These cells have distinct surface markers, control specific signaling pathways, and play an important role in the development of glioma vessels (4,5). Niches have been identified in multiple cancers, and they often assimilate the signals of niches for the tissues from which the CSCs appeared (6,8). However, it is not understood how these cells still manage to preserve their nature when they invade and migrate from their homes to other areas where ideal niches are less likely to be available (9).
Brain tumors and metastases represent a heterogeneous set of conditions (10). Primary brain tumors such as astrocytic tumors, oligodendrogliomas, ependymomas, and mixed gliomas are all referred to as gliomas (11). Adults' most usual lethal brain tumor is grade IV glioma also known as GBM, with over 10,000 cases diagnosed annually in the United States (12). Evidence shows that a population of GBM cells has a stable ability to self-renew and produce new tumors that keep the features of original tumors (13,14).
The ability of stem cells to invade and migrate raises the question of how these cells maintain their stemness when they
come into contact with varied ingredient compositions of a new microenvironment (15). Therefore, looking for genetic changes could help CSCs to keep stemness outside their niches (16). The Quaking gene (QKI), as a tumor suppressor gene, may poten¬tially affect CSC stemness. The gene encodes QKI, a STAR-family RNA-binding protein that is involved in RNA homeostasis (19,20). The RNA-binding protein QKI is highly expresses in brain cells (17), and play a regulatory role in brain development via modulating the stability of mRNAs that promote differentiation and inhibiting cell cycle progress (18). In the present study, the significance of QKI mRNA in human central nervous system (CNS) cancer was evaluated by using the Cancer Genome Atlas (TCGA) data portals.
MATERIALS AND METHODS
Analysis for QKI mutations
The Catalog of Somatic Mutations in Cancer (COSMIC) database was used to analyze QKI mutations (21). Pie charts were generated for a distribution survey and substitutions on the coding strand in CNS cancer.
CBioPortal analysis for alteration frequency of QKI
Alteration frequency of QKI mRNA was performed using cBioPortal for Cancer Genomics (22,23). All searches were performed according to the cBioPortal’s online instructions. The database query was based on mutation and altered expression of the QKI in six CNS/brain studies (Brain Low Grade Glioma (TCGA, Firehose Legacy), Brain Low Grade Glioma (TCGA PanCancer Atlas), GBM (TCGA, Cell 2013), GBM (TCGA, Nature 2008), Low-Grade Gliomas (UCSF, Science 2014), Merged Cohort of LCG and GBM (TCGA, Cell 2016), including 3010 samples of 2958 patients (summarized in Figure 1). The overlapped studies were discarded.

Serial analysis of gene expression (SAGE)
All available published SAGE data were used to analyze QKI expression in normal and cancerous tissues. Digital QKI gene expression profiles were analyzed using SAGE Genie tools (24).
Kaplan-Meier plotter analysis
The prognostic value of the QKI gene in CNS cancer was analyzed using the Kaplan-Meier plotter (25) and PPISURV (26). Overall survival of the patient with high and low levels of QKI was shown by using a Kaplan-Meier survival plot.
RESULTS
QKI mutation in glioma
The information of mutations and mutation types were generated using cBioPortal. Mutation data for QKI were only provided in 17 patients (Table 1). Missense mutation rate was 35.29% (6/17). Furthermore, deletions resulting from frameshifts were observed in 29.41% (5/17) of mutant samples of glioma cancer. Missense mutations wereG163984476 in 50% (3/6), G163956153 in 33.33% (2/6), and C163899861 in 16.67% (1/6) of patients. Alteration frequency of QKI mutation in CNS cancer was analyzed by using cBioPortal. Moreover, 56 of 2,958 (2%) patients had altered QKI gene.
QKI mRNA in Glioma cancer tissues
The expression profile of QKI was found by using the SAGE Digital Gene Expression Display. Higher levels of QKI mRNA were mainly in the brain, spinal cord, breast, stomach, skin, and muscle cancer tissues, compared with their matched normal tissues (Figure 2).
No correlation was observed between the QKI gene expression and its relative protein copy numbers in different types of QKI gene alterations (Figure 3), while the opposite role of QKI mRNA was observed in CNS cancer (p=0.005). In addition, we analyzed the prognostic roles of QKI mRNA in subtypes of CNS cancer, and the results showed that QKI mRNA had no influence on mixed-type CNS cancer (p=0.14).
 Figure 3. The mRNA expression of QKI
Figure 3. The mRNA expression of QKI
All available published SAGE data were used to analyze QKI expression in normal and cancerous tissues. Digital QKI gene expression profiles were analyzed using SAGE Genie tools (24).
Kaplan-Meier plotter analysis
The prognostic value of the QKI gene in CNS cancer was analyzed using the Kaplan-Meier plotter (25) and PPISURV (26). Overall survival of the patient with high and low levels of QKI was shown by using a Kaplan-Meier survival plot.
RESULTS
QKI mutation in glioma
The information of mutations and mutation types were generated using cBioPortal. Mutation data for QKI were only provided in 17 patients (Table 1). Missense mutation rate was 35.29% (6/17). Furthermore, deletions resulting from frameshifts were observed in 29.41% (5/17) of mutant samples of glioma cancer. Missense mutations wereG163984476 in 50% (3/6), G163956153 in 33.33% (2/6), and C163899861 in 16.67% (1/6) of patients. Alteration frequency of QKI mutation in CNS cancer was analyzed by using cBioPortal. Moreover, 56 of 2,958 (2%) patients had altered QKI gene.
QKI mRNA in Glioma cancer tissues
The expression profile of QKI was found by using the SAGE Digital Gene Expression Display. Higher levels of QKI mRNA were mainly in the brain, spinal cord, breast, stomach, skin, and muscle cancer tissues, compared with their matched normal tissues (Figure 2).
No correlation was observed between the QKI gene expression and its relative protein copy numbers in different types of QKI gene alterations (Figure 3), while the opposite role of QKI mRNA was observed in CNS cancer (p=0.005). In addition, we analyzed the prognostic roles of QKI mRNA in subtypes of CNS cancer, and the results showed that QKI mRNA had no influence on mixed-type CNS cancer (p=0.14).
| Sample ID | Cancer type | Cancer Type Detailed | Protein Change | Mutation Type | Copy | Variant Type | Sex |
| TCGA-28-5208-01 | Glioma | Diffuse Glioma | V99Lfs*31 | Frame_Shift_Del | DEL | Diploid | Male |
| TCGA-06-0650-01 | Glioma | Diffuse Glioma | P181R | Missense_Mutation | SNP | Diploid | Female |
| TCGA-06-0155-01 | Glioma | Diffuse Glioma | Q112P | Missense_Mutation | SNP | ShallowDel | Male |
| TCGA-28-5207-01 | Glioma | Diffuse Glioma | R256Kfs*18 | Frame_Shift_Del | DEL | DeepDel | Male |
| TCGA-06-0237-01 | Glioma | Diffuse Glioma | X312_splice | Splice_Site | DEL | Gain | Female |
| TCGA-OX-A56R-01 | Glioma | Diffuse Glioma | E174del | In_Frame_Del | DEL | ShallowDel | Male |
| TCGA-FG-5965-01 | Glioma | Diffuse Glioma | A220G | Missense_Mutation | SNP | Diploid | Female |
| TCGA-FG-5965-01 | Glioma | Oligoastrocytoma | A220G | Missense_Mutation | SNP | Diploid | Female |
| TCGA-FG-5965-01 | Diffuse Glioma | Oligoastrocytoma | A220G | Missense_Mutation | SNP | Diploid | Female |
| TCGA-28-5208-01 | Glioma | Glioblastoma Multiforme | V99Lfs*31 | Frame_Shift_Del | DEL | Diploid | Male |
| TCGA-06-0650-01 | Glioma | Glioblastoma Multiforme | P181R | Missense_Mutation | SNP | Diploid | Female |
| TCGA-28-5207-01 | Glioma | Glioblastoma Multiforme | R256Kfs*18 | Frame_Shift_Del | DEL | DeepDel | Male |
| TCGA-06-0237-01 | Glioma | Glioblastoma Multiforme | X312_splice | Splice_Site | DEL | Gain | Female |
| TCGA-HT-7689-01 | Diffuse Glioma | Oligodendroglioma | QKI-AKAP7 | fusion | FUSION | ShallowDel | Female |
| TCGA-DU-6404-01 | Diffuse Glioma | Oligodendroglioma | PACRG-QKI | fusion | FUSION | Diploid | Female |
| TCGA-HT-8114-01 | Diffuse Glioma | Oligoastrocytoma | PACRG-QKI | fusion | FUSION | Diploid | Male |
| TCGA-WY-A859-01 | Diffuse Glioma | Astrocytoma | NSD2-QKI | fusion | FUSION | Diploid | Female |
Data includes six duplicate mutations in patients with multiple samples
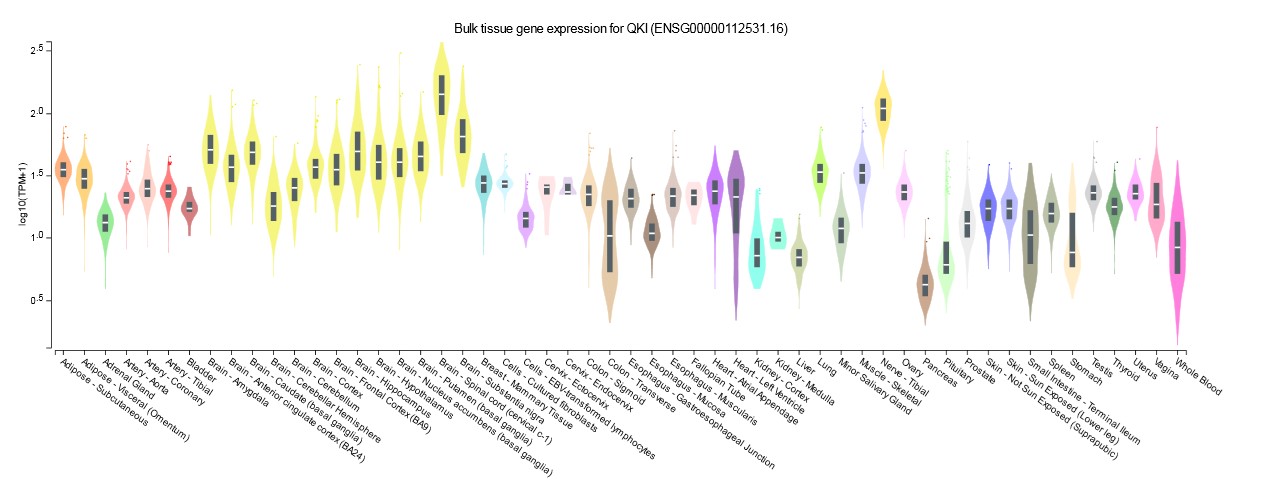
Figure 2. The alteration frequencies of the QKI gene across different cancer studies. Expression of the QKI gene was highest in brain (GTEx portal) (38).

Figure 2. The alteration frequencies of the QKI gene across different cancer studies. Expression of the QKI gene was highest in brain (GTEx portal) (38).

Determination of QKI gene alterations across different cancers and overall survival rate
The QKI gene alterations (including mutations) led to amino acid changes with deleterious impacts on protein function. These included P181R, Q112P, and A220G. The QKI mutations and deletions were less frequent than amplifications in cancer patients. In addition, the QKI gene was notably amplified in several cancers, including stomach, skin, colon, and CNS cancers. In stomach cancer, QKI protein mutations were observed in 17 patients.
Further data showed significantly reduced overall survival rate of patients with altered QKI gene when compared to the unaltered group (Figure 4).
.jpg)
The QKI gene alterations (including mutations) led to amino acid changes with deleterious impacts on protein function. These included P181R, Q112P, and A220G. The QKI mutations and deletions were less frequent than amplifications in cancer patients. In addition, the QKI gene was notably amplified in several cancers, including stomach, skin, colon, and CNS cancers. In stomach cancer, QKI protein mutations were observed in 17 patients.
Further data showed significantly reduced overall survival rate of patients with altered QKI gene when compared to the unaltered group (Figure 4).
.jpg)
DISCUSSION
The niche is unique where preserved stem cells are undifferentiated and circulated through self-renewal. As seen for somatic stem cells, many studies have identified a niche for CSCs that are important for preserving their self-renewal and promoting tumorigenesis (6,8). However, these seemingly specialized CSC niches are absent when CSCs invade other areas or tissues by various routes and metastasize to distant organs via circulation. This raises the question how CSCs keep their self-renewal power outside their niches. Shingu et al. showed that postnatal deletion of Pten and Trp53 could expand the neural stem cell (NSC) population in subventricular zones (SVZs). The lack of NSC self-renewal outside SVZs suggests the inability of NSCs to uphold self-renewal outside their niches, thereby preventing Pten−/−; Trp53−/− premalignant-NSCs from developing into gliomas. They also suggested that Qk deletion in Pten−/−; Trp53−/− premalignant-NSCs improved self-renewal, especially outside their niches, promoting gliomagenesis (3).
Previously, Zheng et al. showed that deletion of Pten and Trp53 in embryonic NSCs could raise sphere-forming capacity/stemness by upregulating Myc, leading to low-grade development and showing that embryonic development NSCs are more manageable to transform than postnatal NSCs (27). Nevertheless, in the model of Zheng et al. (with deletion of Pten and Trp53 in embryonic NSCs), GBM developed with a much more extended latency period (105 days versus 300 days) and lower penetrance (92% versus 25%), suggesting that other genetic or epigenetic changes still must be gained to allow embryonic premalignant neural stem cells to preserve self-renewal outside their niches, to per¬mit full progression of glioma.
The QKI gene has long been studied as a critical gene for oligodendrocyte differentiation and myelin formation (20,28). Other studies confirmed that QKI is needed for oligodendrocyte differentiation (3). This gene is also important for developing smooth muscle, endothelial cells, and monocytes or macrophages (20). In addition, QKI is expressed in NSCs and is a major regulator of self-renewal and differentiation. It has been shown that QKI may regulate RNA homeostasis, including RNA stability, splicing, translation, miRNA processing, and circular RNA biogenesis (19,21).
In support of the importance of QKI as a tumor sup¬pressor, a previous analysis of the TCGA database of GBM demonstrated QKI as the sole gene within the small common region of the 6q26 deletions (32% deletion rate and 1.7% mutation rate) (3,29). Furthermore, QKI downregulation by methylation of the QKI locus (chromosome 6, base 163,755,107) was also reported in 50 of 250 (20%) GBM samples (30). In angiocentric glioma, nearly 90% of tumors have an MYB-QKI translocation, which disrupts QKI (31,32). Other than in gliomas, QKI is a tumor suppressor in other malignancies, including gastric (33), breast (34), colon (35), prostate (36), and oral (37) cancers.
CONCLUSION
Expression of the QKI gene is highest in the brain tissue. Altered QKI gene expression/mutation is correlated with short survival rates in patients with gliomas. The results of the present study suggest that the QKI gene alteration could be considered as a novel prognosis biomarker for CNS/brain cancers.
ACKNOWLEDGMENTS
None.
DECLARATIONS
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethics approvals and consent to participate
Not applicable.
Conflict of interest
The authors declare that there is no conflict of interest regarding publication of this article.
The niche is unique where preserved stem cells are undifferentiated and circulated through self-renewal. As seen for somatic stem cells, many studies have identified a niche for CSCs that are important for preserving their self-renewal and promoting tumorigenesis (6,8). However, these seemingly specialized CSC niches are absent when CSCs invade other areas or tissues by various routes and metastasize to distant organs via circulation. This raises the question how CSCs keep their self-renewal power outside their niches. Shingu et al. showed that postnatal deletion of Pten and Trp53 could expand the neural stem cell (NSC) population in subventricular zones (SVZs). The lack of NSC self-renewal outside SVZs suggests the inability of NSCs to uphold self-renewal outside their niches, thereby preventing Pten−/−; Trp53−/− premalignant-NSCs from developing into gliomas. They also suggested that Qk deletion in Pten−/−; Trp53−/− premalignant-NSCs improved self-renewal, especially outside their niches, promoting gliomagenesis (3).
Previously, Zheng et al. showed that deletion of Pten and Trp53 in embryonic NSCs could raise sphere-forming capacity/stemness by upregulating Myc, leading to low-grade development and showing that embryonic development NSCs are more manageable to transform than postnatal NSCs (27). Nevertheless, in the model of Zheng et al. (with deletion of Pten and Trp53 in embryonic NSCs), GBM developed with a much more extended latency period (105 days versus 300 days) and lower penetrance (92% versus 25%), suggesting that other genetic or epigenetic changes still must be gained to allow embryonic premalignant neural stem cells to preserve self-renewal outside their niches, to per¬mit full progression of glioma.
The QKI gene has long been studied as a critical gene for oligodendrocyte differentiation and myelin formation (20,28). Other studies confirmed that QKI is needed for oligodendrocyte differentiation (3). This gene is also important for developing smooth muscle, endothelial cells, and monocytes or macrophages (20). In addition, QKI is expressed in NSCs and is a major regulator of self-renewal and differentiation. It has been shown that QKI may regulate RNA homeostasis, including RNA stability, splicing, translation, miRNA processing, and circular RNA biogenesis (19,21).
In support of the importance of QKI as a tumor sup¬pressor, a previous analysis of the TCGA database of GBM demonstrated QKI as the sole gene within the small common region of the 6q26 deletions (32% deletion rate and 1.7% mutation rate) (3,29). Furthermore, QKI downregulation by methylation of the QKI locus (chromosome 6, base 163,755,107) was also reported in 50 of 250 (20%) GBM samples (30). In angiocentric glioma, nearly 90% of tumors have an MYB-QKI translocation, which disrupts QKI (31,32). Other than in gliomas, QKI is a tumor suppressor in other malignancies, including gastric (33), breast (34), colon (35), prostate (36), and oral (37) cancers.
CONCLUSION
Expression of the QKI gene is highest in the brain tissue. Altered QKI gene expression/mutation is correlated with short survival rates in patients with gliomas. The results of the present study suggest that the QKI gene alteration could be considered as a novel prognosis biomarker for CNS/brain cancers.
ACKNOWLEDGMENTS
None.
DECLARATIONS
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethics approvals and consent to participate
Not applicable.
Conflict of interest
The authors declare that there is no conflict of interest regarding publication of this article.
Article Type: Research |
Subject:
Cellular and Molecular Biology
References
1. López de Andrés J, Griñán-Lisón C, Jiménez G, Marchal JA. Cancer stem cell secretome in the tumor microenvironment: a key point for an effective personalized cancer treatment. Journal of hematology & oncology. 2020 Dec;13(1):1-22. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
2. Shenghui HE, Nakada D, Morrison SJ. Mechanisms of stem cell self-renewal. Annual Review of Cell and Developmental. 2009 Nov 10;25:377-406. [View at Publisher] [DOI] [PMID] [Google Scholar]
3. Shingu T, Ho AL, Yuan L, Zhou X, Dai C, Zheng S, Wang Q, Zhong Y, Chang Q, Horner JW, Liebelt BD. Qki deficiency maintains stemness of glioma stem cells in suboptimal environment by downregulating endolysosomal degradation. Nature genetics. 2017 Jan;49(1):75-86. [View at Publisher] [Google Scholar]
4. Liebelt BD, Shingu T, Zhou X, Ren J, Shin SA, Hu J. Glioma stem cells: signaling, microenvironment, and therapy. Stem cells international. 2016 Oct;2016. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
5. Alves AL, Gomes IN, Carloni AC, Rosa MN, da Silva LS, Evangelista AF, Reis RM, Silva VA. Role of glioblastoma stem cells in cancer therapeutic resistance: a perspective on antineoplastic agents from natural sources and chemical derivatives. Stem Cell Research & Therapy. 2021 Dec;12(1):1-22. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
6. Calabrese C, Poppleton H, Kocak M, Hogg TL, Fuller C, Hamner B, Oh EY, Gaber MW, Finklestein D, Allen M, Frank A. A perivascular niche for brain tumor stem cells. Cancer cell. 2007 Jan 1;11(1):69-82. [View at Publisher] [Google Scholar]
7. Li Z, Bao S, Wu Q, Wang H, Eyler C, Sathornsumetee S, Shi Q, Cao Y, Lathia J, McLendon RE, Hjelmeland AB. Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer cell. 2009 Jun 2;15(6):501-13. [View at Publisher] [Google Scholar]
8. Merlos-Suárez A, Barriga FM, Jung P, Iglesias M, Céspedes MV, Rossell D, Sevillano M, Hernando-Momblona X, da Silva-Diz V, Muñoz P, Clevers H. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell stem cell. 2011 May 6;8(5):511-24. [View at Publisher] [Google Scholar]
9. Li L, Neaves WB. Normal stem cells and cancer stem cells: the niche matters. Cancer research. 2006 May 1;66(9):4553-7. [View at Publisher] [DOI] [PMID] [Google Scholar]
10. Focusing on brain tumours and brain metastasis. Nat Rev Cancer. 2019 Dec ;20(1):1-1. [View at Publisher] [DOI] [PMID]
11. Hanif F, Muzaffar K, Perveen K, Malhi SM, Simjee SU. Glioblastoma multiforme: a review of its epidemiology and pathogenesis through clinical presentation and treatment. Asian Pacific journal of cancer prevention: APJCP. 2017;18(1):3. [PubMed] [Google Scholar]
12. Dunn GP, Rinne ML, Wykosky J, Genovese G, Quayle SN, Dunn IF, Agarwalla PK, Chheda MG, Campos B, Wang A, Brennan C. Emerging insights into the molecular and cellular basis of glioblastoma. Genes & development. 2012 Apr 15;26(8):756-84. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
13. Rich JN, Eyler CE. Cancer stem cells in brain tumor biology. InCold Spring Harbor symposia on quantitative biology 2008 Jan 1 (Vol. 73, pp. 411-420). Cold Spring Harbor Laboratory Press. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
14. Vescovi AL, Galli R, Reynolds BA. Brain tumour stem cells. Nature Reviews Cancer. 2006 Jun;6(6):425-36. [View at Publisher] [DOI] [PMID] [Google Scholar]
15. Giese A, Bjerkvig R, Berens ME, Westphal M. Cost of migration: invasion of malignant gliomas and implications for treatment. Journal of clinical oncology. 2003 Apr 15;21(8):1624-36. [DOI] [PMID] [Google Scholar]
16. Hu J, Ho AL, Yuan L, Hu B, Hua S, Hwang SS, Zhang J, Hu T, Zheng H, Gan B, Wu G. Neutralization of terminal differentiation in gliomagenesis. Proceedings of the National Academy of Sciences. 2013 Sep 3;110(36):14520-7. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
17. Suiko T, Kobayashi K, Aono K, Kawashima T, Inoue K, Ku L, Feng Y, Koike C. Expression of quaking RNA-binding protein in the adult and developing mouse retina. PLoS One. 2016 May 19;11(5):e0156033. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
18. Sakers K, Liu Y, Llaci L, Lee SM, Vasek MJ, Rieger MA, Brophy S, Tycksen E, Lewis R, Maloney SE, Dougherty JD. Loss of Quaking RNA binding protein disrupts the expression of genes associated with astrocyte maturation in mouse brain. Nature communications. 2021 Mar 9;12(1):1-4. [View at Publisher] [Google Scholar]
19. Chen AJ, Paik JH, Zhang H, Shukla SA, Mortensen R, Hu J, Ying H, Hu B, Hurt J, Farny N, Dong C. STAR RNA-binding protein Quaking suppresses cancer via stabilization of specific miRNA. Genes & development. 2012 Jul 1;26(13):1459-72. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
20. Darbelli L, Richard S. Emerging functions of the Quaking RNA‐binding proteins and link to human diseases. Wiley Interdisciplinary Reviews: RNA. 2016 May;7(3):399-412. [View at Publisher] [DOI] [PMID] [Google Scholar]
21. Forbes S, Bhamra G, Bamford S, Dawson E, Kok C, Clements J, Menzies A, Teague JW, Futreal PA, Stratton MR. The catalogue of somatic mutations in cancer (COSMIC). Current protocols in human genetics. 2008 Apr;57(1):10-1. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
22. Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Science signaling. 2013 Apr 2;6(269):pl1-. [View at Publisher] [DOI] [Google Scholar]
23. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer discovery. 2012 May 1;2(5):401-4. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
24. Velculescu VE, Zhang L, Vogelstein B, Kinzler KW. Serial analysis of gene expression. Science. 1995 Oct 20;270(5235):484-7. [View at Publisher] [DOI] [PMID] [Google Scholar]
25. Szász AM, Lánczky A, Nagy Á, Förster S, Hark K, Green JE, Boussioutas A, Busuttil R, Szabó A, Győrffy B. Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1,065 patients. Oncotarget. 2016 Aug 2;7(31):49322. [DOI] [PMID] [PMCID] [Google Scholar]
26. Antonov AV, Krestyaninova M, Knight RA, Rodchenkov I, Melino G, Barlev NA. PPISURV: a novel bioinformatics tool for uncovering the hidden role of specific genes in cancer survival outcome. Oncogene. 2014 Mar;33(13):1621-8. [View at Publisher] [Google Scholar]
27. Zheng H, Ying H, Yan H, Kimmelman AC, Hiller DJ, Chen AJ, Perry SR, Tonon G, Chu GC, Ding Z, Stommel JM. p53 and Pten control neural and glioma stem/progenitor cell renewal and differentiation. Nature. 2008 Oct;455(7216):1129-33. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
28. Chénard CA, Richard S. New implications for the QUAKING RNA binding protein in human disease. Journal of neuroscience research. 2008 Feb 1;86(2):233-42. [View at Publisher] [DOI] [PMID] [Google Scholar]
29. Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R. The somatic genomic landscape of glioblastoma. Cell. 2014 Apr 24;157(3):753. [Google Scholar]
30. Fuchs E, Tumbar T, Guasch G. Socializing with the neighbors: stem cells and their niche. Cell. 2004 Mar 19;116(6):769-78. [View at Publisher] [Google Scholar]
31. Qaddoumi I, Orisme W, Wen J, Santiago T, Gupta K, Dalton JD, Tang B, Haupfear K, Punchihewa C, Easton J, Mulder H. Genetic alterations in uncommon low-grade neuroepithelial tumors: BRAF, FGFR1, and MYB mutations occur at high frequency and align with morphology. Acta neuropathologica. 2016 Jun;131(6):833-45. [View at Publisher] [Google Scholar]
32. Bandopadhayay P, Ramkissoon LA, Jain P, Bergthold G, Wala J, Zeid R, et al. MYB-QKI rearrangements in angiocentric glioma drive tumorigenicity through a tripartite mechanism. Nat Genet . 2016 Feb; 48(3):273-82 [View at Publisher]
33. Bian Y, Wang L, Lu H, Yang G, Zhang Z, Fu H, Lu X, Wei M, Sun J, Zhao Q, Dong G. Downregulation of tumor suppressor QKI in gastric cancer and its implication in cancer prognosis. Biochemical and biophysical research communications. 2012 May 25;422(1):187-93. [View at Publisher] [Google Scholar]
34. Yu F, Jin L, Yang G, Ji L, Wang F, Lu Z. Post-transcriptional repression of FOXO1 by QKI results in low levels of FOXO1 expression in breast cancer cells. Oncology reports. 2014 Mar 1;31(3):1459-65. [View at Publisher] [DOI] [PMID] [Google Scholar]
35. Yang G, Fu H, Zhang J, Lu X, Yu F, Jin L, Bai L, Huang B, Shen L, Feng Y, Yao L. RNA-binding protein quaking, a critical regulator of colon epithelial differentiation and a suppressor of colon cancer. Gastroenterology. 2010 Jan 1;138(1):231-40. [View at Publisher] [Google Scholar]
36. Zhao YI, Zhang G, Wei M, Lu X, Fu H, Feng F, Wang S, Lu W, Wu N, Lu Z, Yuan J. The tumor suppressing effects of QKI-5 in prostate cancer: a novel diagnostic and prognostic protein. Cancer biology & therapy. 2014 Jan 1;15(1):108-18. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
37. Lu W, Feng F, Xu J, Lu X, Wang S, Wang L, Lu H, Wei M, Yang G, Wang L, Lu Z. QKI impairs self-renewal and tumorigenicity of oral cancer cells via repression of SOX2. Cancer biology & therapy. 2014 Sep 1;15(9):1174-84. [View at Publisher] [DOI] [PMID] [PMCID] [Google Scholar]
38. Lonsdale J, Thomas J, Salvatore M, Phillips R, Lo E, Shad S, Hasz R, Walters G, Garcia F, Young N, Foster B. The genotype-tissue expression (GTEx) project. Nature genetics. 2013 Jun;45(6):580-5. [View at Publisher] [Google Scholar]
Send email to the article author
| Rights and permissions | |
 |
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License. |

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International (CC BY-NC 4.0).





